

Flexible tool for bias detection, visualization, and mitigation. Use
models explained with DALEX and calculate
fairness classification metrics based on confusion matrices using
fairness_check() or try newly developed module for
regression models using fairness_check_regression(). R
package fairmodels allows to compare and gain information about various
machine learning models. Mitigate bias with various pre-processing and
post-processing techniques. Make sure your models are classifying
protected groups similarly.
Install it from CRAN:
install.packages("fairmodels")or developer version from GitHub:
devtools::install_github("ModelOriented/fairmodels")Checking fairness is easy!
library(fairmodels)
library(ranger)
library(DALEX)
data("german")
# ------------ step 1 - create model(s) -----------------
lm_model <- glm(Risk~.,
data = german,
family=binomial(link="logit"))
rf_model <- ranger(Risk ~.,
data = german,
probability = TRUE,
num.trees = 200)
# ------------ step 2 - create explainer(s) ------------
# numeric y for explain function
y_numeric <- as.numeric(german$Risk) -1
explainer_lm <- explain(lm_model, data = german[,-1], y = y_numeric)
explainer_rf <- explain(rf_model, data = german[,-1], y = y_numeric)
# ------------ step 3 - fairness check -----------------
fobject <- fairness_check(explainer_lm, explainer_rf,
protected = german$Sex,
privileged = "male")
print(fobject)
plot(fobject)
Compas recidivism data use case: Basic
tutorial
Bias mitigation techniques on Adult data: Advanced
tutorial
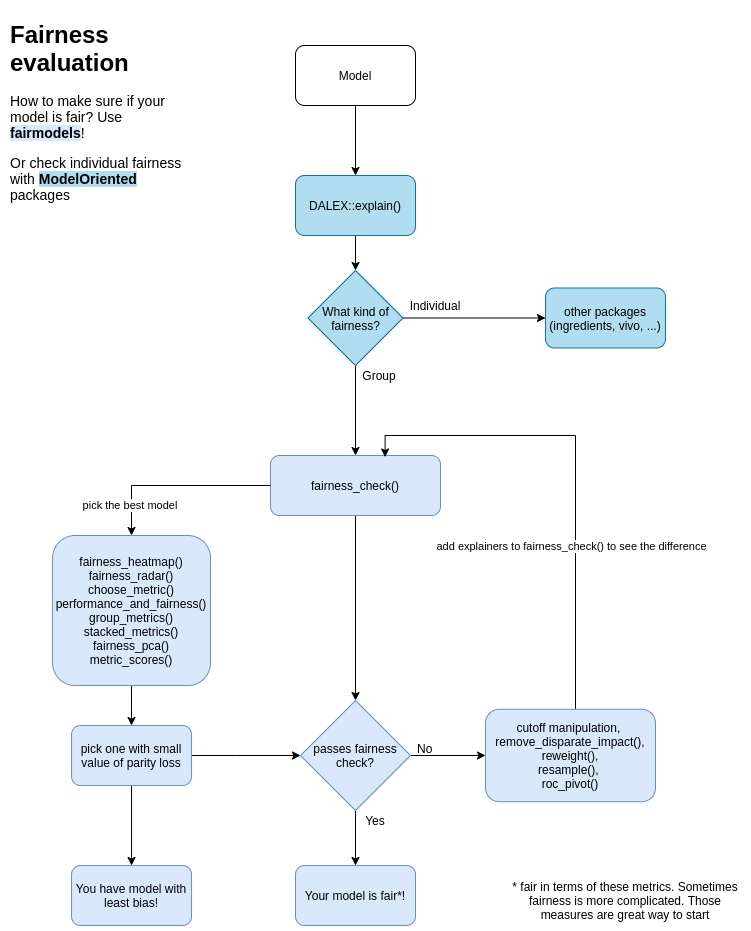
fairness_check parameters are
* x, … - explainers and fairness_objects
(products of fairness_check).
* protected - factor with different subgroups as levels. Usually
specific race, sex etc…
* privileged - subgroup, base on which to calculate parity loss
metrics.
* cutoff - custom cutoff, might be single value - cutoff same for all
subgroups or vector - for each subgroup individually. Affecting only
explainers.
* label - character vector for every explainer.
Models might be trained on different data, even without protected
variable. May have different cutoffs which gives different values of
metrics. fairness_check() is place where
explainers and fairness_objects are checked
for compatibility and then glued together.
So it is possible to to something like this:
fairness_object <- fairness_check(explainer1, explainer2, ...)
fairness_object <- fairness_check(explainer3, explainer4, fairness_object, ...)even with more fairness_objects!
If one is even more keen to know how fairmodels works
and what are relations between objects, please look at this diagram class
diagram
There are 12 metrics based on confusion matrix :
| Metric | Formula | Full name | fairness names while checking among subgroups |
|---|---|---|---|
| TPR | 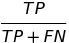 |
true positive rate | equal opportunity |
| TNR | 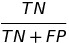 |
true negative rate | |
| PPV | 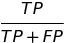 |
positive predictive value | predictive parity |
| NPV | 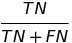 |
negative predictive value | |
| FNR | 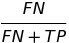 |
false negative rate | |
| FPR | 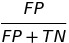 |
false positive rate | predictive equality |
| FDR | 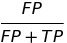 |
false discovery rate | |
| FOR | 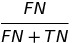 |
false omission rate | |
| TS | 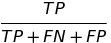 |
threat score | |
| STP | 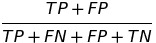 |
statistical parity | statistical parity |
| ACC | 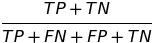 |
accuracy | Overall accuracy equality |
| F1 |  |
F1 score |
and their parity loss.
How is parity loss calculated?
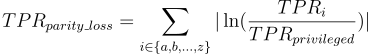
Where i denotes the membership to unique subgroup from
protected variable. Unprivileged subgroups are represented by small
letters and privileged by simply “privileged”.
some fairness metrics like Equalized odds are satisfied if parity loss in both TPR and FPR is low
It is relatively easy! Check it out here
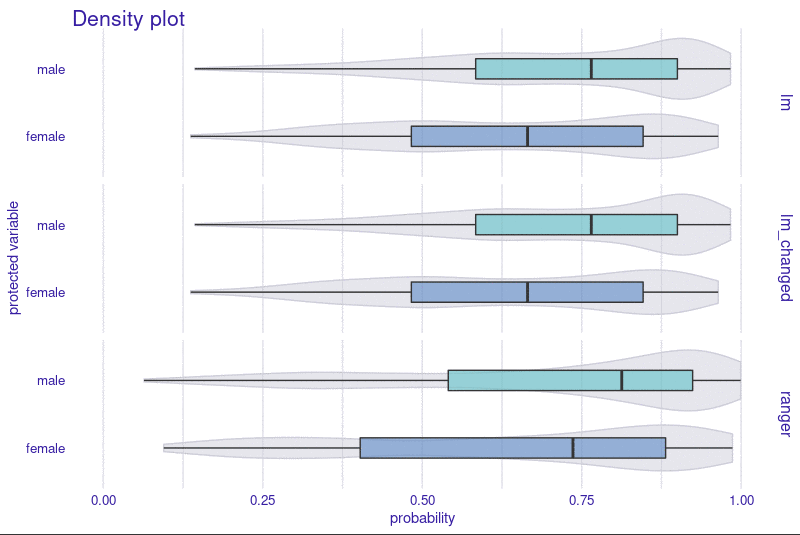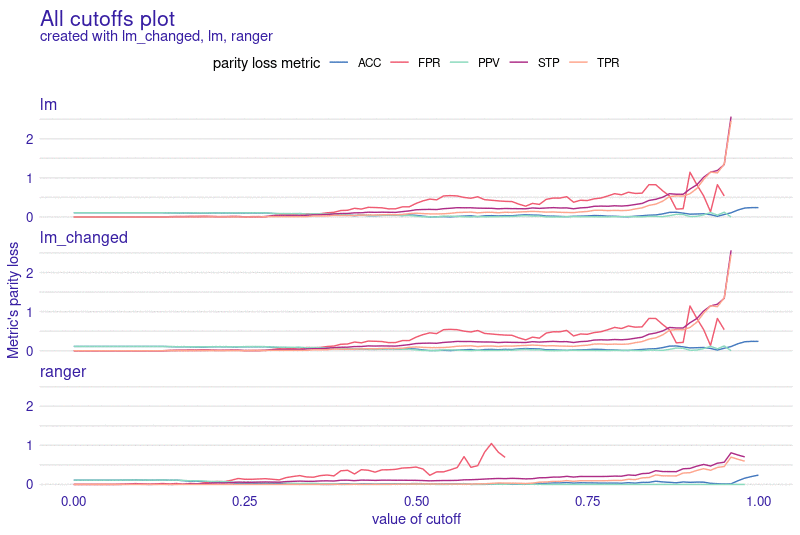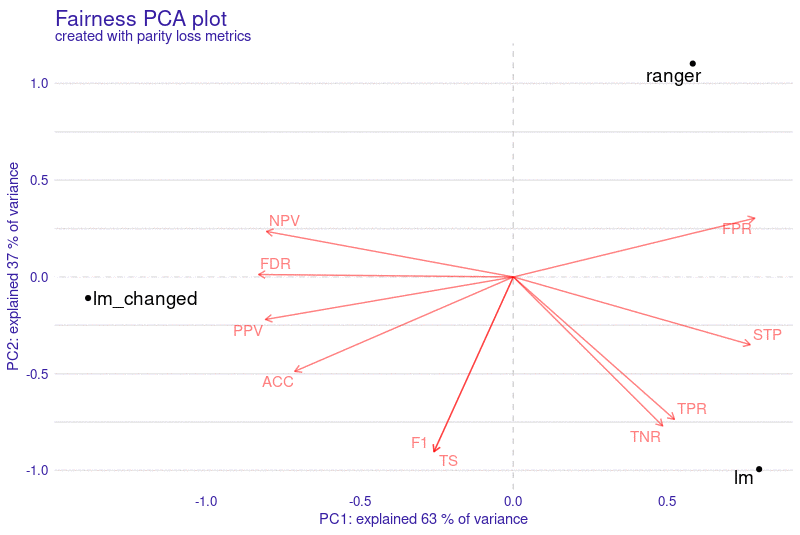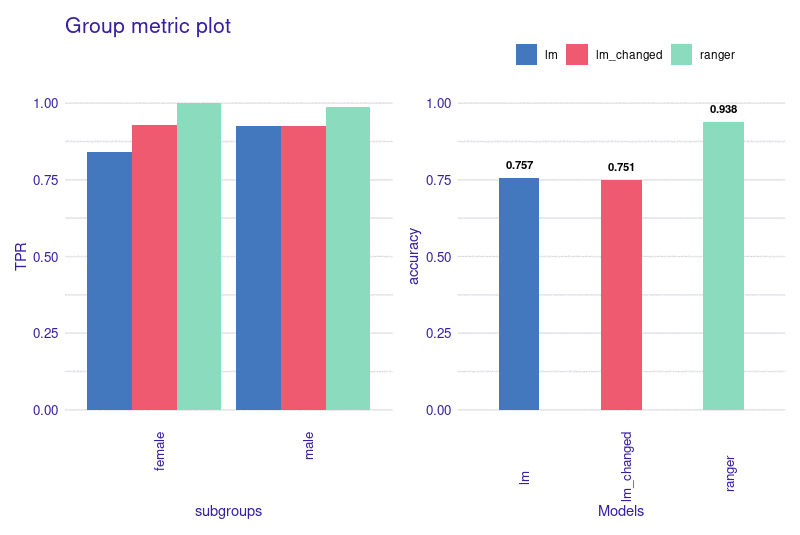
R package fairmodels has support for regression models. Check
fairness using fairness_check_regression() to approximate
classification fairness metrics in regression setting. Plot object with
plot() to visualize fairness check or with
plot_density() to see model’s output.
Zafar, Valera, Rodriguez, Gummadi (2017) https://arxiv.org/pdf/1610.08452.pdf
Barocas, Hardt, Narayanan (2019) https://fairmlbook.org/
Steinberg, Daniel & Reid, Alistair & O’Callaghan, Simon. (2020). Fairness Measures for Regression via Probabilistic Classification. - https://arxiv.org/pdf/2001.06089.pdf