The goal of nlmixr2plot is to provide the nlmixr2 core estimation routines.
You can install the development version of nlmixr2plot from GitHub with:
# install.packages("remotes")
remotes::install_github("nlmixr2/nlmixr2data")
remotes::install_github("nlmixr2/lotri")
remotes::install_github("nlmixr2/rxode2")
remotes::install_github("nlmixr2/nlmixr2est")
remotes::install_github("nlmixr2/nlmixr2extra")
remotes::install_github("nlmixr2/nlmixr2plot")For most people, using nlmixr2 directly would be likely easier.
library(nlmixr2est)
#> Loading required package: nlmixr2data
library(nlmixr2plot)
## The basic model consists of an ini block that has initial estimates
one.compartment <- function() {
ini({
tka <- 0.45 ; label("Log Ka")
tcl <- 1 ; label("Log Cl")
tv <- 3.45 ; label("Log V")
eta.ka ~ 0.6
eta.cl ~ 0.3
eta.v ~ 0.1
add.sd <- 0.7
})
# and a model block with the error specification and model specification
model({
ka <- exp(tka + eta.ka)
cl <- exp(tcl + eta.cl)
v <- exp(tv + eta.v)
d/dt(depot) = -ka * depot
d/dt(center) = ka * depot - cl / v * center
cp = center / v
cp ~ add(add.sd)
})
}
## The fit is performed by the function nlmixr/nlmix2 specifying the model, data and estimate
fit <- nlmixr2(one.compartment, theo_sd, est="saem", saemControl(print=0))
#> → loading into symengine environment...
#> → pruning branches (`if`/`else`) of saem model...
#> ✔ done
#> → finding duplicate expressions in saem model...
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#> → optimizing duplicate expressions in saem model...
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#> ✔ done
#> rxode2 2.0.11.9000 using 8 threads (see ?getRxThreads)
#> no cache: create with `rxCreateCache()`
#> Calculating covariance matrix
#> → loading into symengine environment...
#> → pruning branches (`if`/`else`) of saem model...
#> ✔ done
#> → finding duplicate expressions in saem predOnly model 0...
#> → finding duplicate expressions in saem predOnly model 1...
#> → optimizing duplicate expressions in saem predOnly model 1...
#> → finding duplicate expressions in saem predOnly model 2...
#> ✔ done
#> → Calculating residuals/tables
#> ✔ done
#> → compress origData in nlmixr2 object, save 5952
#> → compress phiM in nlmixr2 object, save 62360
#> → compress parHist in nlmixr2 object, save 9592
#> → compress saem0 in nlmixr2 object, save 28760
print(fit)
#> ── nlmixr² SAEM OBJF by FOCEi approximation ──
#>
#> Gaussian/Laplacian Likelihoods: AIC() or $objf etc.
#> FOCEi CWRES & Likelihoods: addCwres()
#>
#> ── Time (sec $time): ──
#>
#> setup covariance saem table compress other
#> elapsed 0.002 0.01 5.24 0.1 0.09 2.498
#>
#> ── Population Parameters ($parFixed or $parFixedDf): ──
#>
#> Parameter Est. SE %RSE Back-transformed(95%CI) BSV(CV%) Shrink(SD)%
#> tka Log Ka 0.454 0.196 43.1 1.57 (1.07, 2.31) 71.5 -0.0203%
#> tcl Log Cl 1.02 0.0853 8.4 2.76 (2.34, 3.26) 27.6 3.46%
#> tv Log V 3.45 0.0454 1.32 31.5 (28.8, 34.4) 13.4 9.89%
#> add.sd 0.693 0.693
#>
#> Covariance Type ($covMethod): linFim
#> No correlations in between subject variability (BSV) matrix
#> Full BSV covariance ($omega) or correlation ($omegaR; diagonals=SDs)
#> Distribution stats (mean/skewness/kurtosis/p-value) available in $shrink
#> Censoring ($censInformation): No censoring
#>
#> ── Fit Data (object is a modified tibble): ──
#> # A tibble: 132 × 19
#> ID TIME DV PRED RES IPRED IRES IWRES eta.ka eta.cl eta.v cp
#> <fct> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 1 0 0.74 0 0.74 0 0.74 1.07 0.103 -0.491 -0.0820 0
#> 2 1 0.25 2.84 3.27 -0.426 3.87 -1.03 -1.48 0.103 -0.491 -0.0820 3.87
#> 3 1 0.57 6.57 5.85 0.723 6.82 -0.246 -0.356 0.103 -0.491 -0.0820 6.82
#> # … with 129 more rows, and 7 more variables: depot <dbl>, center <dbl>,
#> # ka <dbl>, cl <dbl>, v <dbl>, tad <dbl>, dosenum <dbl>
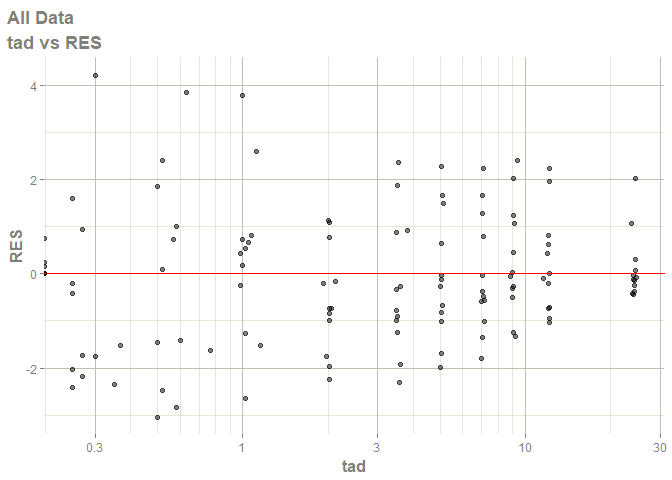
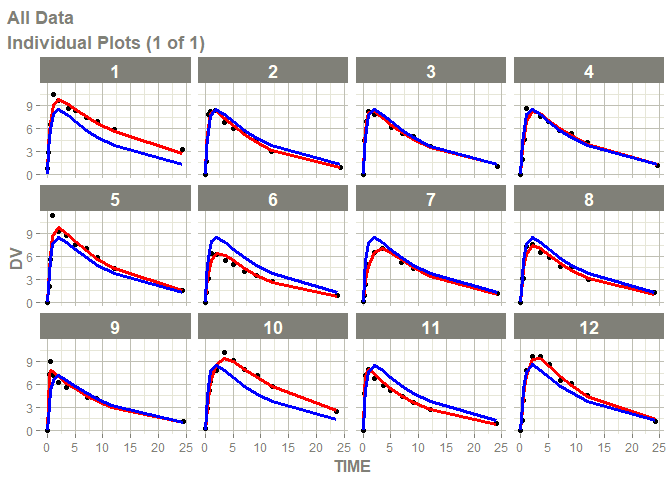
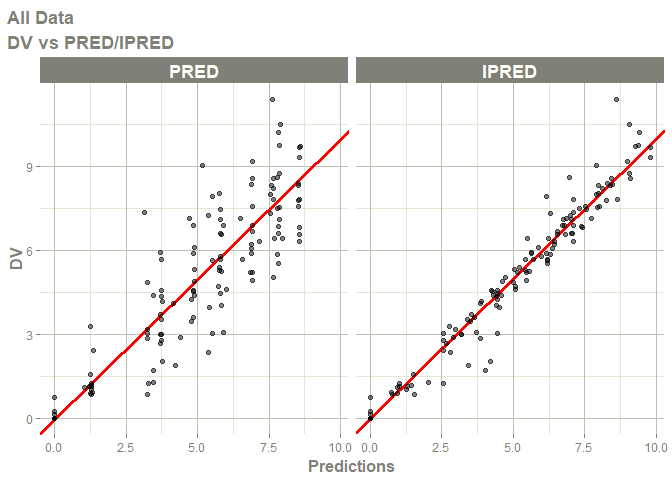
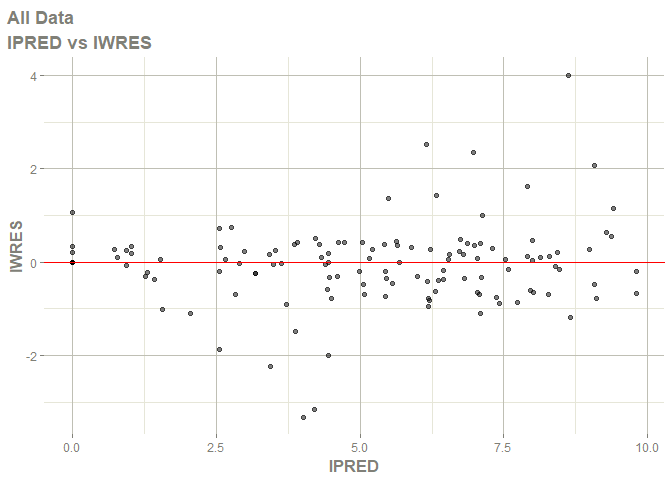
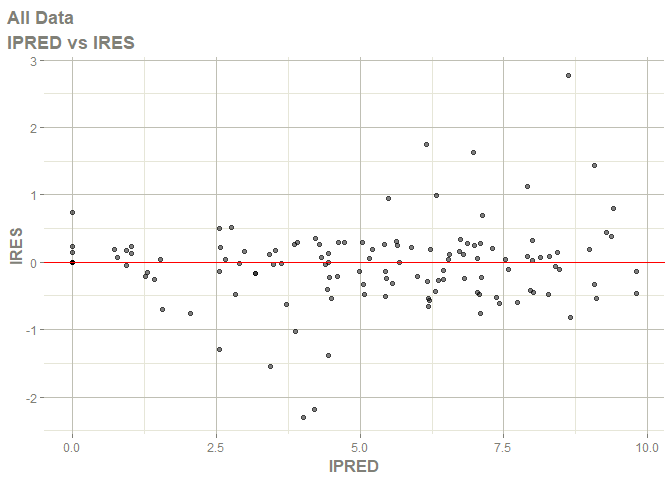
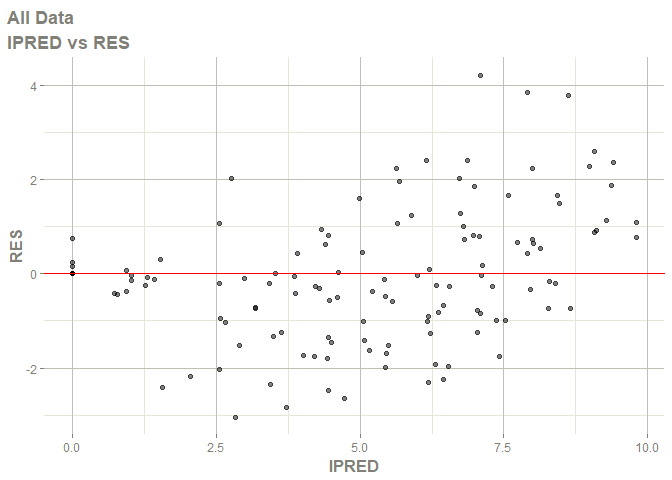
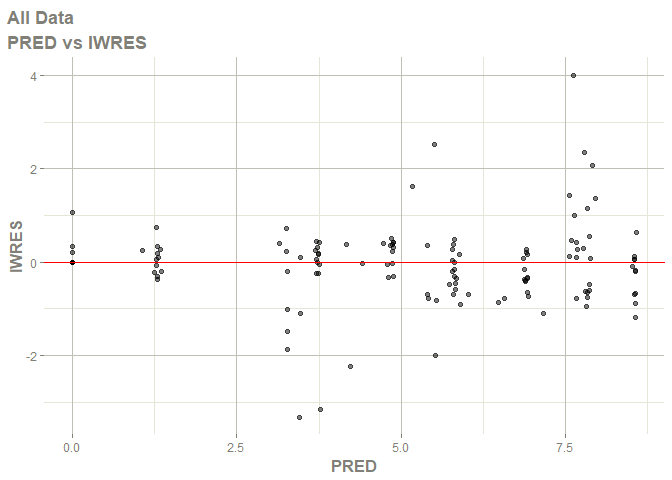
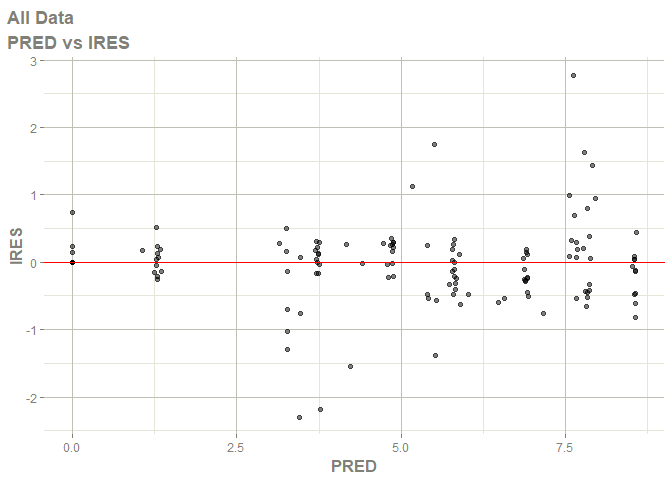
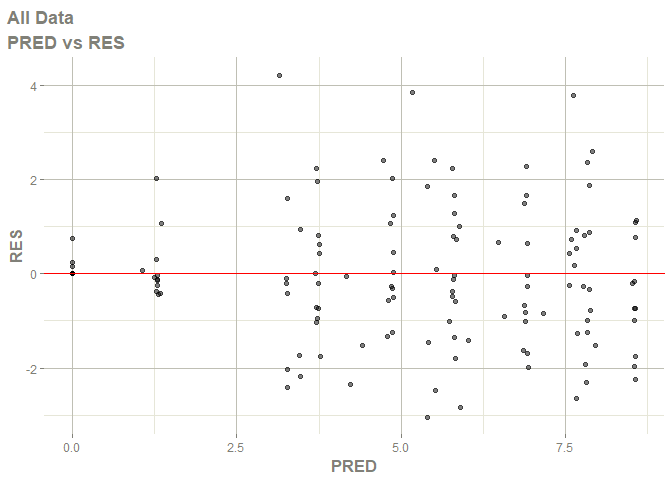
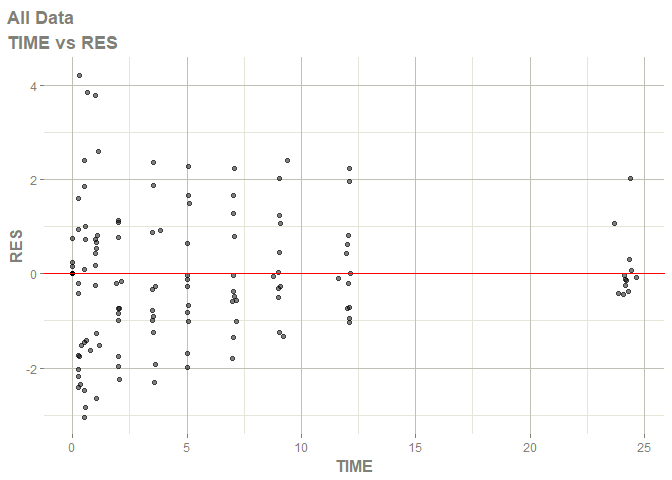
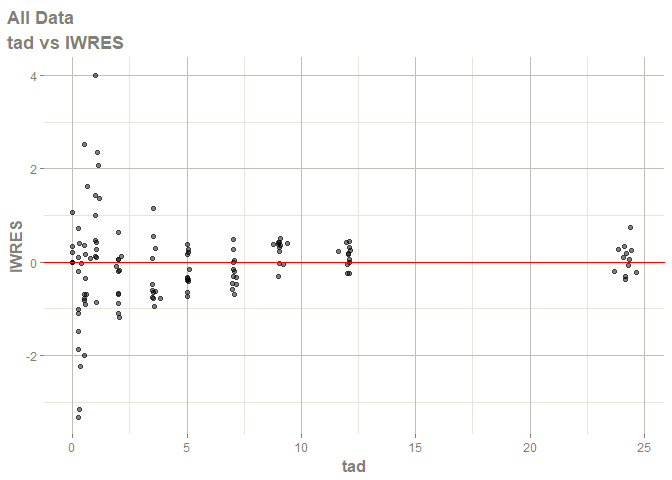
# this now gives the goodness of fit plots
plot(fit)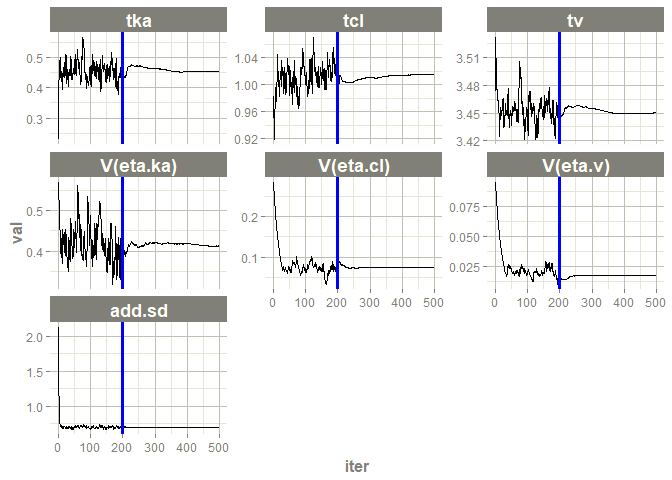
#> Warning: Transformation introduced infinite values in continuous x-axis
#> Warning: Transformation introduced infinite values in continuous y-axis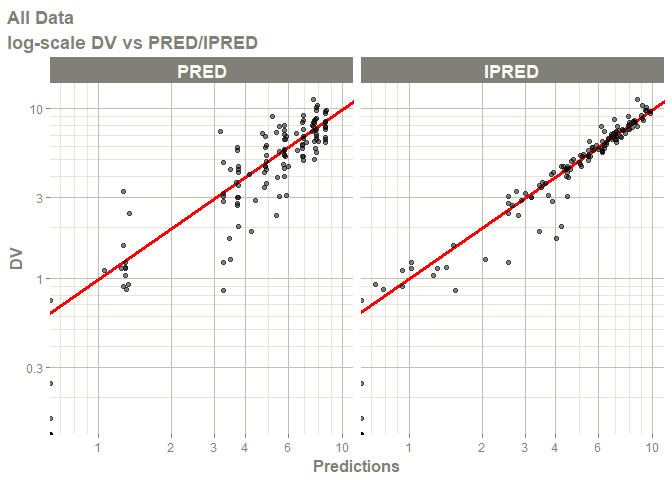
#> Warning: Transformation introduced infinite values in continuous x-axis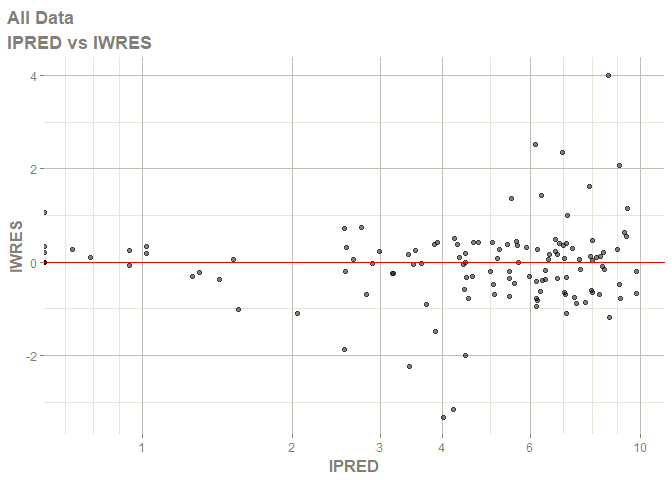
#> Warning: Transformation introduced infinite values in continuous x-axis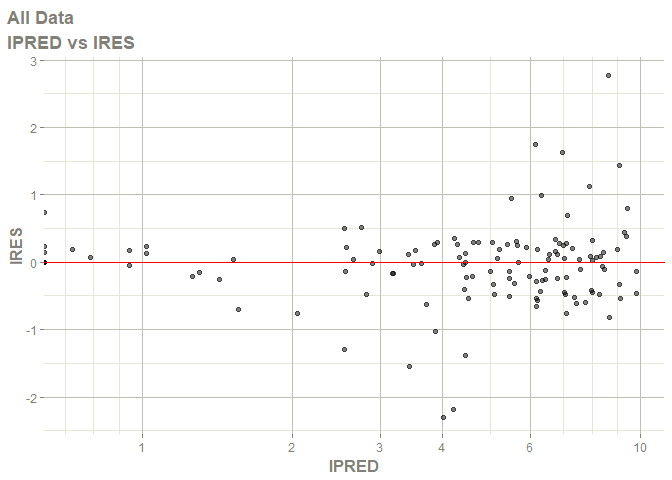
#> Warning: Transformation introduced infinite values in continuous x-axis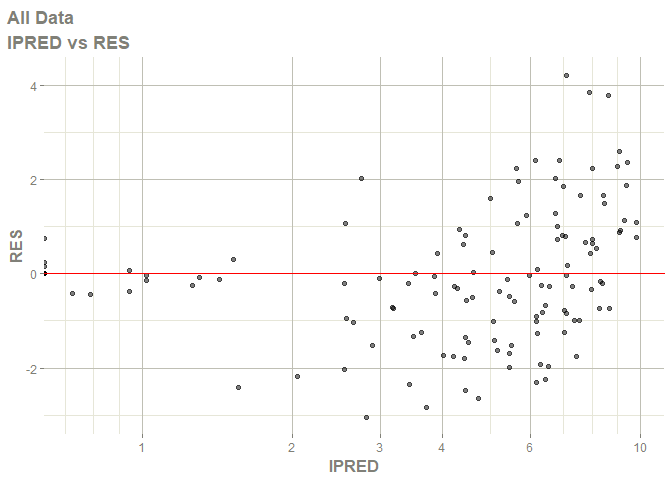
#> Warning: Transformation introduced infinite values in continuous x-axis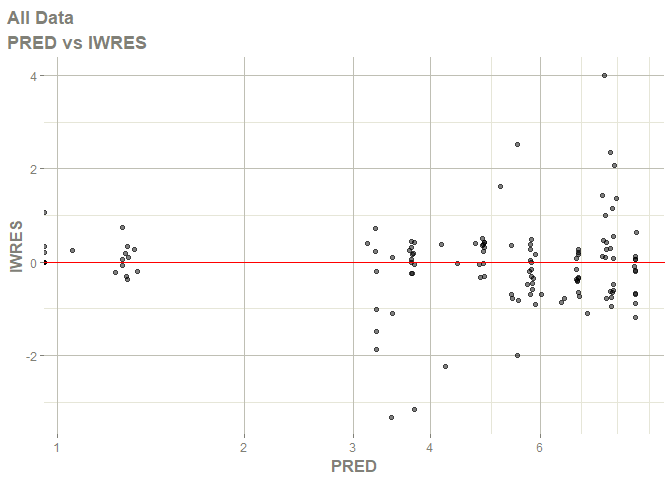
#> Warning: Transformation introduced infinite values in continuous x-axis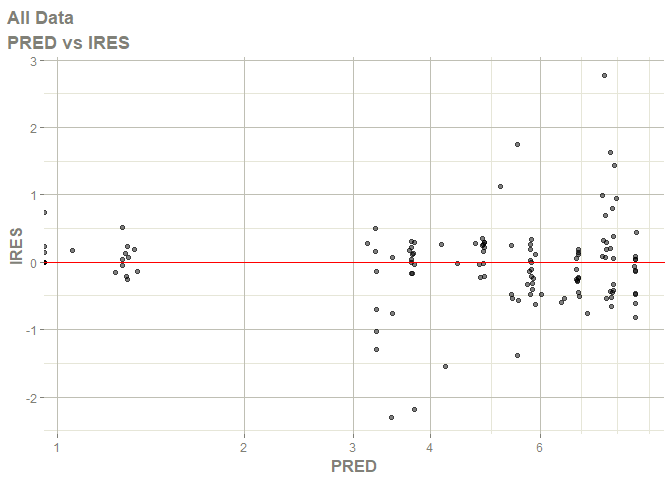
#> Warning: Transformation introduced infinite values in continuous x-axis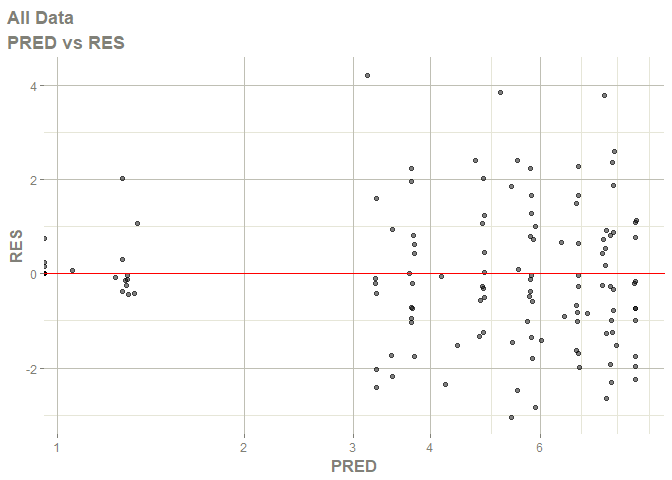
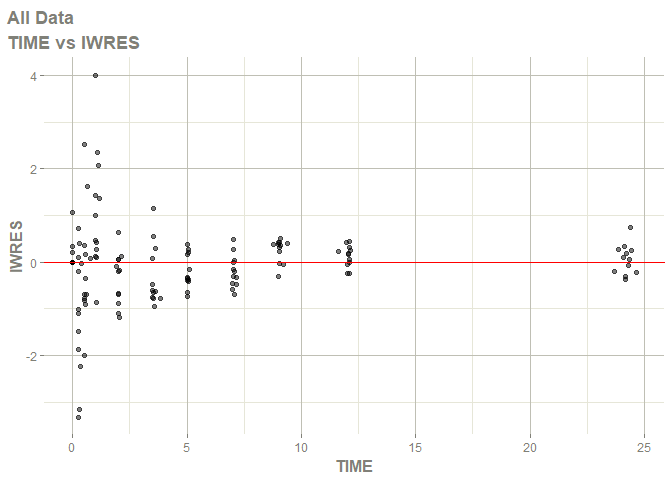
#> Warning: Transformation introduced infinite values in continuous x-axis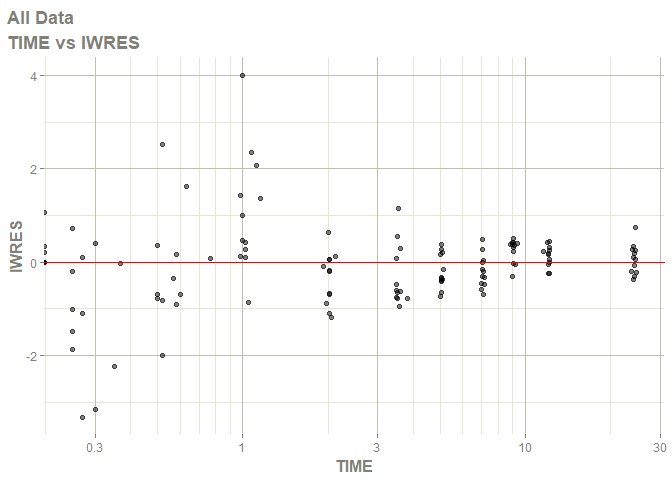
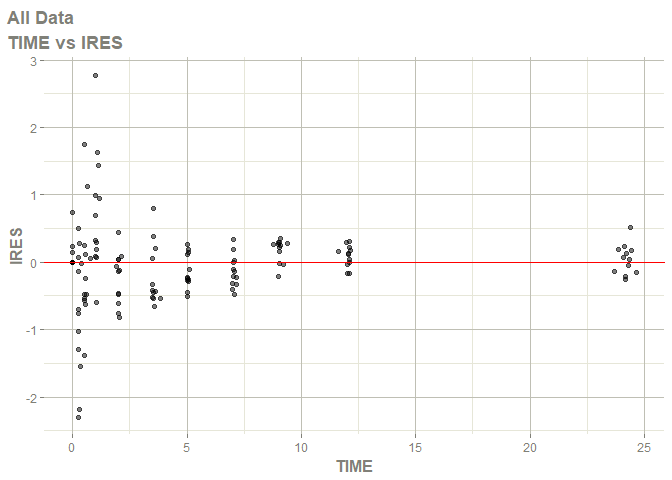
#> Warning: Transformation introduced infinite values in continuous x-axis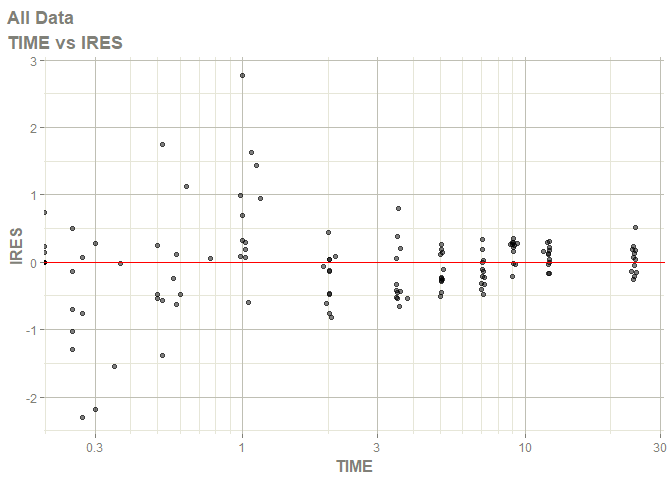
#> Warning: Transformation introduced infinite values in continuous x-axis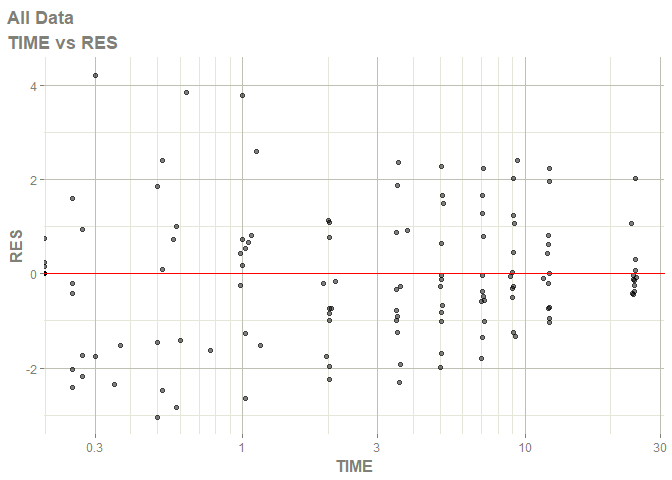
#> Warning: Transformation introduced infinite values in continuous x-axis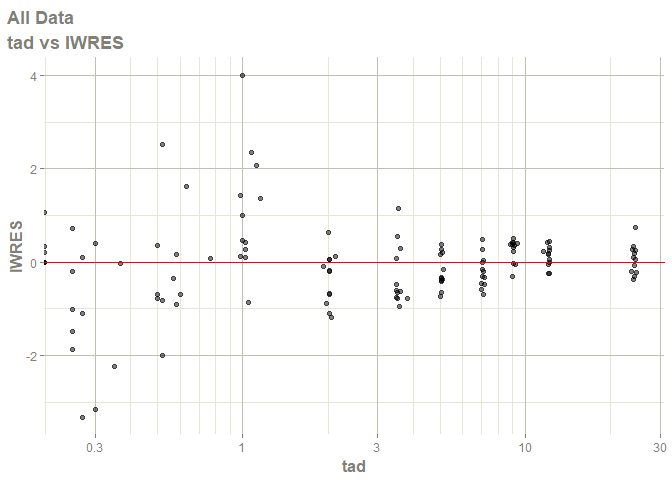
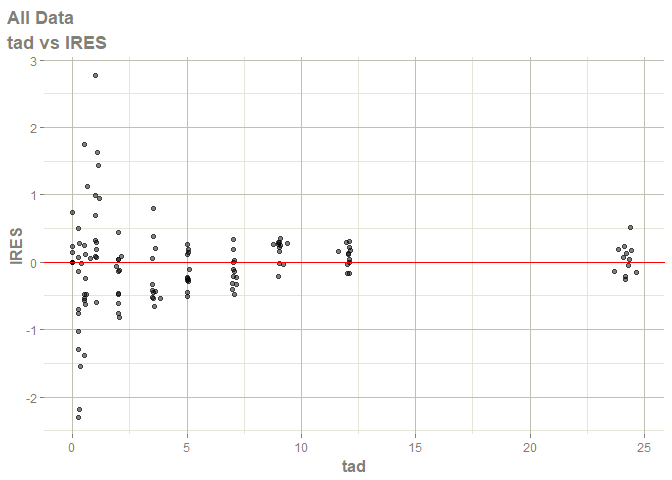
#> Warning: Transformation introduced infinite values in continuous x-axis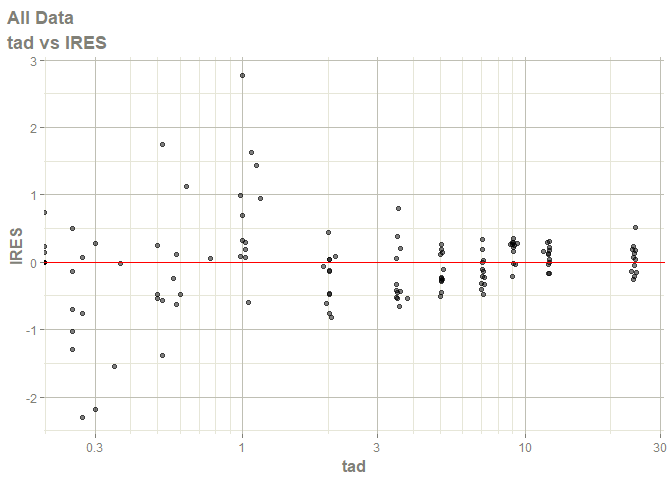
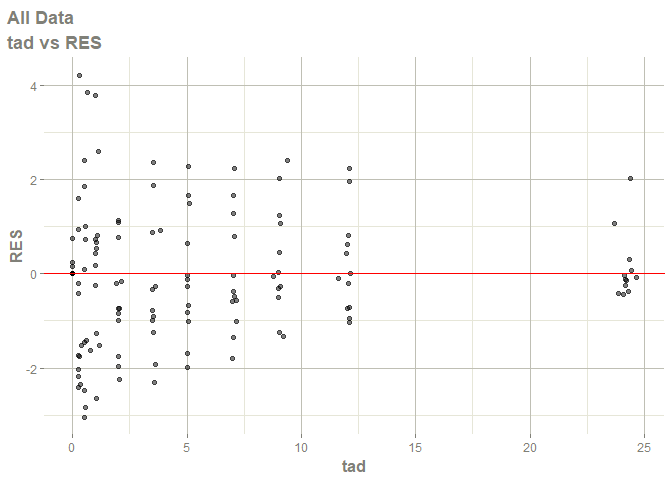
#> Warning: Transformation introduced infinite values in continuous x-axis